Outreach activities
The Aegean Archipelago: a living laboratory of evolutionary biology
Introduction
Through funding provided by the European Society of Evolutionary Biology we developed outreach material for primary schools on the Aegean Archipelago which is a living laboratory of evolutionary biology.
The material is available in Greek and English at present
The material comprises three games:
- A board game entitled "Giants and Dwarfs"
- A card game entitled "Aegean’s Settlers"
- An online game entitled "Aerial Collisions"
The goal of the games is to
- explain to children that the Earth was not always as we know it today, and that life forms on Earth change over time.
- our planet has been and will be for a long time to come, but life on it is changing; species go extinct, and others are taking their place and evolving, in an endless cycle of speciation and extinction.
- via the online game, children become familiar with the concept of DNA and the genetic code, based on which the characteristics of each organism are formed. The more similar the DNA of two organisms is, the more closely related they will be (in general, we know that there are exceptions).
Download Area and Teacher preparation material
- GREEK
- ENGLISH
-
AERIAL COLLISIONS GAME: 10 minute preparatory videos for teachers in English and Greek
The team behind the games
The games were designed by scientists from the Natural History Museum of Crete, the Karlsruhe Institute of Technology, the Heidelberg Institute for Theoretical Studies, the Carnegie Institute for Science in Stanford, and the Technical University of Munich.
We also wish to thank the nice wildlife team of FraPort Greece for their kind support.
Pilot deployment phase
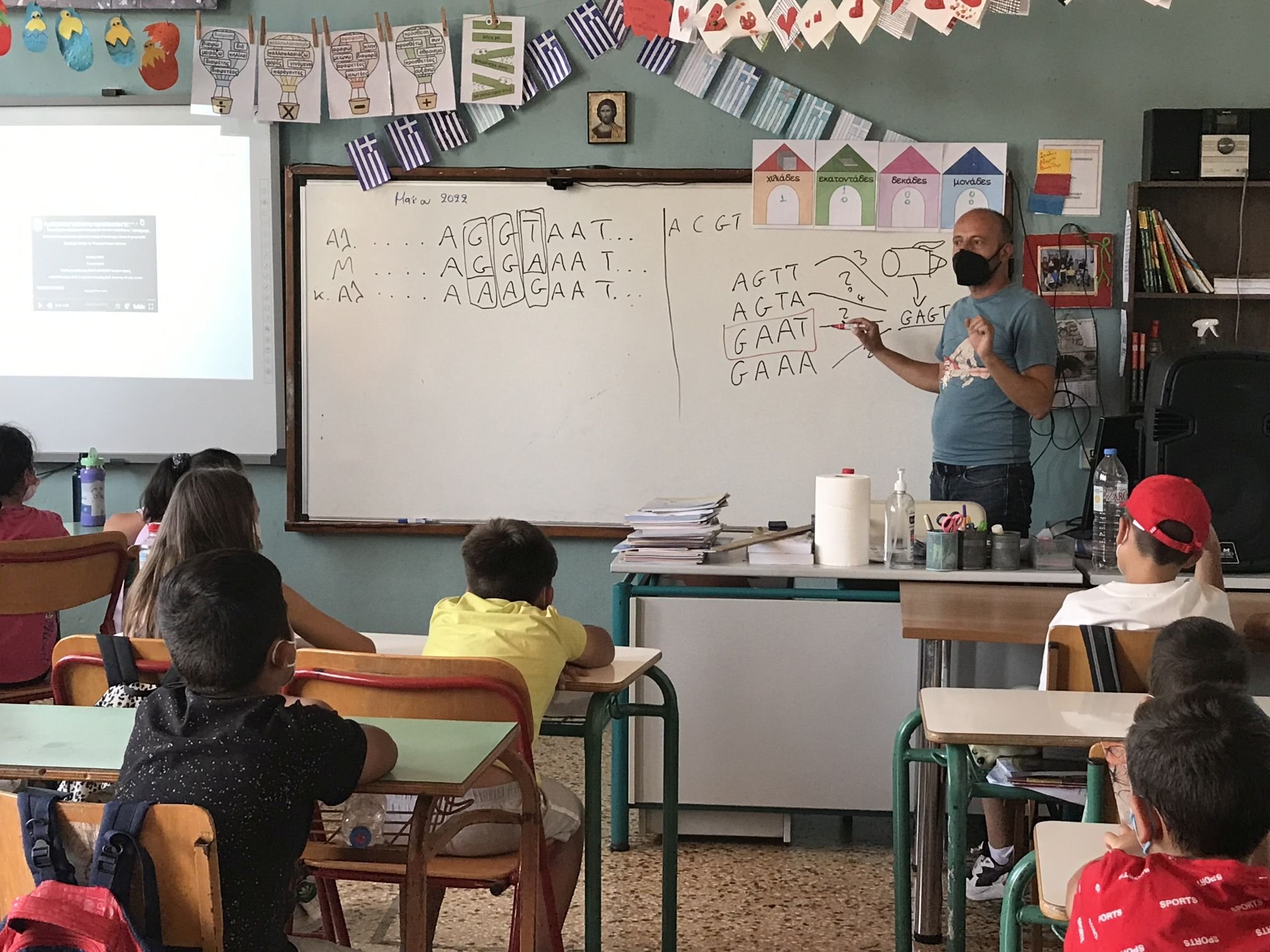
The initial testing phase in schools around Crete, with an emphasis on remote and rural schools, was completed by end of May 2022. The material is available now to the scientific/educational community (in Greek and English).
The image below shows Alexis presenting the material at the primary school of Sivas in southern Crete.

Videos
In July 2022 Alexis gave an outreach talk at HITS about using phylogenetic methods for predicting elimination tournaments ins sports.
In a 2021 online conversation, Casey Dunn, Professor of Ecology and Evolutionary Biology at Yale University, and Alexis Stamatakis talk about the RAxML source code, discuss the tension between generality and optimization when developing software, and explore the interface between software and engineering. You can watch the whole conversation on YouTube:
A TV report in English about our recent paper in Science that was the first outcome of the 1KITE project.
A TV report by the nano technology magazine of 3Sat about the work of our group in the framework of the 1KITE project.
Alexis gave a video interview (in German) about his new duties as professor of computer science at KIT in conjunction with his position at HITS.
Local Cretan TV (Kriti TV) report about 2012 summer school on computational molecular evolution sponsored by EMBO and co-sponsored by HITS.
Local Cretan TV (Kriti TV) report about 2010 summer school on computational molecular evolution sponsored by EMBO:
Agniezska Borowska from HITS produced a very nice video about phylogenetic inference: